On March 6, 2019, Douglas E. Soltis (Distinguished Professor in the Laboratory of Molecular Systematics & Evolutionary Genetics at the Florida Museum of Natural History and the Department of Biology at the University of Florida) and Pamela S. Soltis (Distinguished Professor and Curator in the Florida Museum of Natural History at the University of Florida and Director of the University of Florida Biodiversity Institute) spoke at the Academy about a project that harnesses algorithm development, computer power, and DNA sequencing to create a comprehensive visual Tree of Life. The program, which served as the 2079th Stated Meeting of the Academy, included a welcome from David W. Oxtoby (President of the Academy) and an introduction from Scott Vernon Edwards (Alexander Agassiz Professor of Zoology at Harvard University and Curator of Ornithology at the Museum of Comparative Zoology). An abridged version of Douglas Soltis’s and Pamela Soltis’s presentations appears below.

What we would like to do this evening is to take you on a journey with the Tree of Life. To start, what do we mean when we say the Tree of Life? Well, really, we are talking about biodiversity: the totality of life on Earth – all 2.3 million described species and perhaps one hundred million species that have yet to be discovered, described, and named. A. E. Waite really sums up the importance of the metaphor of the Tree of Life throughout human history: “Behind the Man is the Tree of Life.” Throughout human history, cultures have used the image of a tree to represent their connectivity to all living things. If you imagine a giant tree with leaves: we are but one leaf with all the other species on our planet that are connected to that tree.
The Tree of Life imagery has had spiritual, religious, magical, and mystical connotations to people throughout human history. One might say that the Tree of Life is really a concept that is as old as our own species. There are examples of Tree of Life artifacts from around the world that date back as far as 4000 B.C. The Tree of Life is mentioned over ten times in the Bible, but it is usually the Tree of Knowledge of Good and Evil that gets all the attention.
The Mayan Tree of Life is similar to many other trees of life, connecting species as well as gods. Connections to the underworld run through the roots of the tree. Furthermore, in many of these Tree of Life stories, damage to the tree was taboo. The Hopi Indians in the Southwest of the United States, as part of their Tree of Life legend, refer to an earlier civilization that abused and overexploited the Tree of Life, which ultimately led to their demise. Words to live by today!
The modern concept of the Tree of Life traces to the time of Darwin. As he said in On the Origin of Species: “and so by generation I believe it has been with the great Tree of Life, which fills with its dead and broken branches the crust of the earth, and covers the surface with its ever-branching and beautiful ramifications.”
The Tree of Life idea was so important to Darwin that the only diagram that he included in On the Origin of Species is his sketch of the Tree of Life. It is a diagram that shows a tree of relationships. This concept inspired other biologists of the time, including the famous German biologist, Ernst Haeckel, to begin to draw and depict what the Tree of Life might look like. And scientists did this for the next 150 years. They drew trees by hand to depict evolutionary relationships, of how they thought organisms were related. It really wasn’t until quite recently, until 2015, that the first comprehensive Tree of Life for all 2.3 million species that have been named so far finally appeared (see Figure 1).
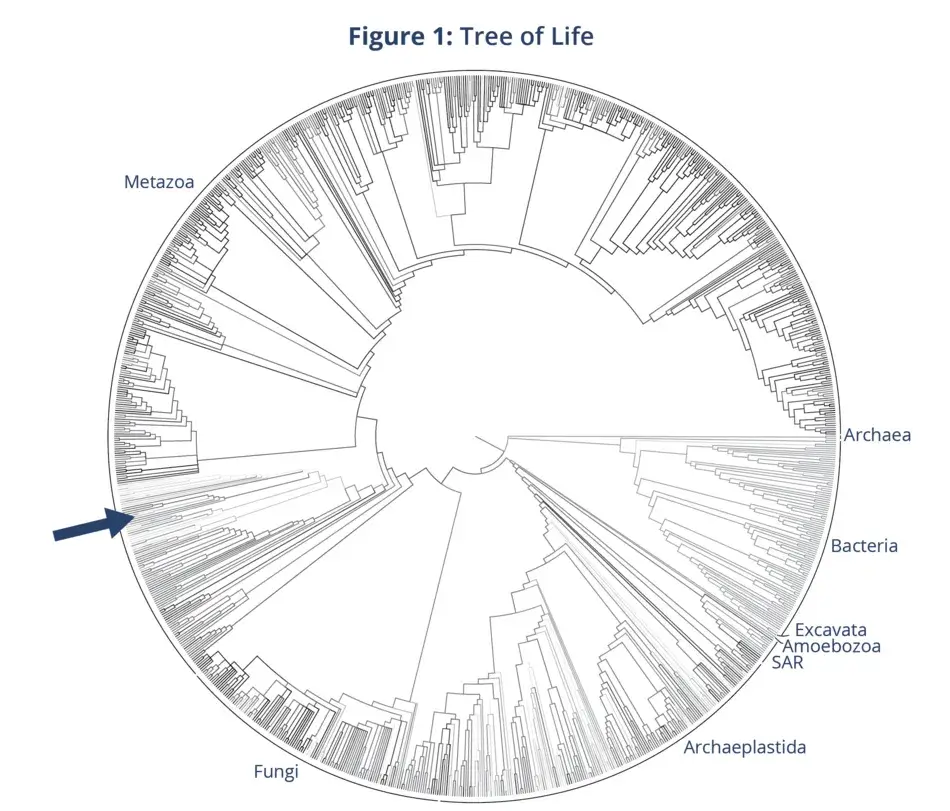
The major groups are labeled around the tree. All 2.3 million species are represented at the tips. The nodes that run throughout the tree are the inferred ancestors. The arrow in the image indicates you; it is the position of Homo sapiens on the Tree of Life. I like these big trees because they are humbling. They show that we are just one small branch on this giant Tree of Life. We don’t get a gold star; we don’t have a larger or thicker branch; we are just one small branch. This is a hypothesis of relationships, a rough draft of what we think the Tree of Life might look like.
Now, if Darwin were here today, he might ask what took so long. There are several reasons why it took so long to build the Tree of Life. The first is that unlike other fields, such as chemistry, physics, and astronomy, tree-building is a very young field. It was only in the 1960s that people started building trees in a somewhat rigorous manner that used data and repeatable methods. And just to make things more confusing, scientists depict trees in various ways. Tree thinking is a recent development when you consider the history of science.
The second reason it took so long is that building trees, especially really big trees, is hard. It rivals some of the most complex problems in mathematics, astronomy, and physics. Let me explain why.
If we start with three species and we try to build a tree of how they are related, there is only one way the species can be connected: three species give us one possible (unrooted) tree. If we have five species, we have fifteen possible trees. With ten species, we have over two million possible trees. As we see, these numbers are increasing logarithmically. By the time you get to twenty species, you are now approaching 1 mole of trees: in other words, 6.02 x 1023 trees. And by the time you get to about two hundred species, the number of possible trees exceeds the number of atoms that we think are in the universe. So, these are big, complex problems. It is no wonder then that building the Tree of Life was long considered a grand challenge in biology, a moonshot for biodiversity science. In fact, there are published papers in esteemed journals including Science that stated it was basically impossible to build a comprehensive Tree of Life.
But we ultimately did it and by we, I mean a large community of scientists. This was a team effort that really reflects how modern scientists operate and how science works. It was a perfect storm of algorithm development, computational power, and DNA sequence data.
Now, just how big is the Tree of Life? How big is a tree that has 2.3 million species? If you printed the Tree of Life as we now know it in a linear format, using standard sheets of paper that measure 8.5 x 11 inches, with species in twelve point font, it would take four sides of fourteen Empire State Buildings just to display the Tree of Life for 2.3 million species.
But we have a long way still to go. We are just at the starting point. As the famous biodiversity scientist E. O. Wilson has said, “It is entirely possible that specialists have discovered only 20 percent, or fewer, of Earth’s biodiversity at the species level. . . . Scientists . . . are in a race to find as many of the surviving species as possible . . . before they vanish and thus are not only overlooked but never to be known.”
Let’s consider our tree again. We have DNA data for only 17 percent of the species on the tree. That is abysmal. We estimate that there are at least ten million more species on our planet that have never been named and have never been included in any tree of relationships. For anybody who is interested in the Tree of Life, there is plenty of work still to do. Now what this doesn’t reflect adequately is the bacterial world. When we consider bacteria, there may be one hundred million species or more that remain unnamed and not placed in any tree. Madonna’s well-known song lyrics, “we are living in a material world,” are off by one word. We are living in a bacterial world!
Even in areas of the world that are well-studied, for example, much of North America, there remain cryptic species that have never been noted, never been identified, because they are difficult to distinguish. Species that look similar to us may be very distinct genetically. And this is very common in the plant, animal, and fungal worlds. What is the value of knowing more about the Tree of Life? Why is it so important? As the prominent geneticist and evolutionary biologist Theodosius Dobzhansky said, “Nothing in biology makes sense except in light of evolution.” The modern corollary to that statement is, “Everything in biology makes more sense in light of a tree of relationships.” We all understand that because of the importance of our own family trees. If you have an ancestor who has a disease that is genetically controlled, you know there is a strong probability that you inherited that trait. Trees of relationships are predictive, whether they are your own family tree or the Tree of Life.
Let’s consider some examples of why it is so important to our well-being to know more about the Tree of Life. First is drug discovery. Most of our medicines come from plants. So how do we find the next generation of medicines? There are about half a million green plant species, so it would take a long time to survey each one individually. That had been the traditional approach. A better approach is to use the tree of relationships, the Tree of Life, and take advantage of what we already know. So, for example, if you are going to look for the next generation of heart medicines, or medicines for nervous disorders, there are only a couple of places in the Tree of Life where you would probably want to look because that is where the plants that yield these compounds have all been found so far. And in those parts of the tree, there are thousands of species that have never been tested for their chemistry. But we know they have those compounds (see Figure 2).
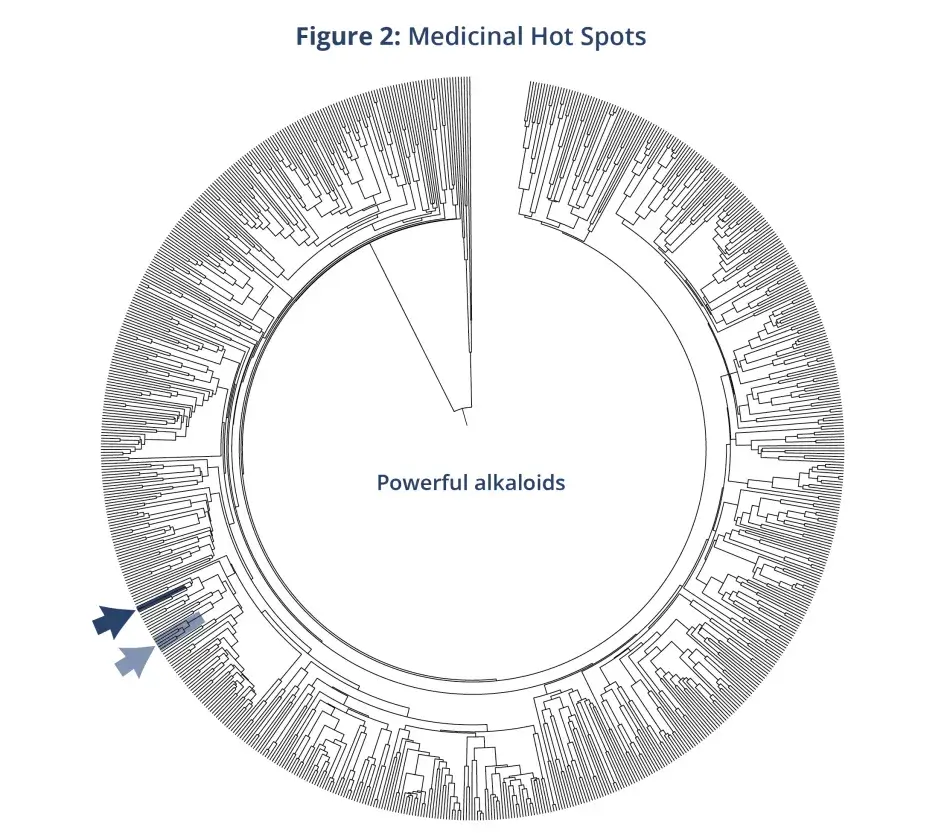
A second example is crop improvement. Where do we obtain genes for disease resistance, water use efficiency, and increased yield? You look for the closest relatives of crops by using the Tree of Life. The following quote from J. D. Miller at the U.S. Department of Agriculture sums up the whole story: “If no germplasm from wild relatives had been used, there would probably not be a viable sugarcane industry in any place in the world.” This is true for many of our crops. Let me share an example from our own lab. We had a student who studied domesticated squashes, things like pumpkin, acorn squash, butternut squash, and the like. If you have ever grown squash, you know that they require a lot of water. So how do we build a better squash? Our student, Heather Rose Kates, built a tree of relationships for squash and their relatives, and she found that the closest relatives of these plants, the ones that we domesticated, are dry-adapted species that are found in the arid Southwest. The collected germplasm can be used in breeding programs to help build a squash that is less water-loving and more dry-adapted.
My third example relates to disease. One of the first lines of defense in studying disease is to build a phylogeny, a tree of relationships. Many of you may remember the SARS epidemic from 2002 – 2003. It started in China; there were more than 750 deaths in thirty-seven countries. It was considered a possible pandemic. But what was the source of SARS? By building a tree of relationships, it became clear that the jump to humans occurred either from civets, a small mammal, or from a species of bat. Trees of relationships are critical in studying disease.
Trees of relationships are also important in the study of conservation, which is my fourth example. There is a little bird called the white-winged warbler. When you do a phylogenetic study and build a tree, you find that this bird is not a true warbler. Furthermore, this bird is found on only one island, the island of Hispaniola. So, by using the Tree of Life, the white-winged warbler became a species of conservation concern. And there are other examples of how using the Tree of Life is important for conservation.
A fifth example is the response of organisms to climate change. Because trees of relationships are predictive, we can use the Tree of Life to help inform which species may be most susceptible to a changing climate. In other words, we can use a big data approach across large parts of the Tree of Life to better inform people as to which organisms, which parts of the Tree of Life, might be most susceptible to a rapidly changing climate, either increasing temperature or a decrease in moisture and other factors.

I would like to continue with this theme of conservation and response to climate change by considering how the Tree of Life can aid conservation efforts. I will be using examples of plants found in Florida. Now, Florida is home to more than four thousand species of plants, many of which occur in very specialized habitats that are already being affected by climate change. When we examine biodiversity hotspots, such as those in California, the Appalachians, and Florida, how do we determine which species or areas to protect? Often decisions are based on the total number of species that are present, or on a so-called indicator species. But what if we used the power of the Tree of Life to help prioritize these efforts and aid conservation decisions? We can do this by using something called phylogenetic diversity, which takes into account the relationships that we see in the Tree of Life. Instead of counting the number of species in an area, we can use their relationships to prioritize conservation decisions.
So how does this work? Well, phylogenetic diversity measures how much of the Tree of Life is present in a given area. Let me use a simple example to explain how it works.
Say we have two areas. The first area has eight species, and they are all oaks. Now to an oak specialist this might seem like great diversity because these oak species may seem different. But to most of us and in relation to the rest of the Tree of Life, these eight species of oaks are all very similar and represent just a very small part of the tree. The second area also has eight species, but they are much more diverse and represent a larger portion of the Tree of Life. This second area would have higher phylogenetic diversity and might be a better target area for conservation because more of the Tree of Life is represented. So, for any area, such as a small region of the Florida Panhandle, we can examine the numbers of species and the types of species that are in that area. We can plot them on a phylogeny and count the amount of branch length, the amount of the tree that is present in the area. To do this, we need a phylogeny or a tree of relationships, which we have built for the flora of Florida. We can get locality information from herbarium specimens, such as the pressed plants that are housed in a museum’s herbarium. We have about a half-million or so of these specimens at the University of Florida in the Florida Museum’s herbarium. Harvard has several million specimens in its herbarium. Or we can go to online repositories such as iDigBio, which is the national center for digitized biodiversity collections, housed at the University of Florida. With a tree of relationships and locality information, we can estimate phylogenetic diversity for a range of locations and plot them on a map, which can then be used to target how we want to set up conservation areas.
We can also compare these patterns of phylogenetic diversity with other aspects of the environment or with information on human populations. This can help inform areas that are of high phylogenetic diversity and of greatest concern in terms of human impact.
How well do our current conservation areas capture phylogenetic diversity? We do not always have the best match. Regions that are preserved may have low phylogenetic diversity, but there are other important reasons for conserving an area. Conversely, areas of high phylogenetic diversity may have only limited conserved space. Future decisions about conservation might be better if they take these phylogenetic diversity measures into account.
Despite our efforts at conservation, species are disappearing rapidly. As Daniel Kozlovsky said, “What your parents can hardly remember, you will not miss. What you now take for granted, or what is slowly disappearing, your children, not having known, cannot lament.” This is a moving statement. Think about it now in relation to the organisms and plants that our parents were familiar with, such as the chestnut tree, that we barely have any association with, or the things that we recognize as being important in our lives that our children or our grandchildren may never have the opportunity to encounter.
So, this brings us to the topic of extinction. We are now in the age referred to by many as the Anthropocene, with extinction rates one thousand times higher than typical and between one hundred and two hundred species being lost per day. These levels of extinction are equivalent to the great mass extinctions, such as the one that included the loss of non-avian dinosaurs. What can we do? Species are being extirpated at alarming rates, and we are continually trying to save those that are endangered. E. O. Wilson has suggested that we set aside half of our planet with sites selected from around the globe in order to preserve at least key regions. This argument certainly has some merit. As it turns out, only about 17 percent of the Earth’s area is set aside in preserves. But any sort of large-scale action requires education. H. G. Wells wrote, “Civilization is a race between education and catastrophe.” Or, if you are from our era, you might remember this chorus from a song by Crosby, Stills, Nash & Young, “Teach your children well . . . and feed them on your dreams.”
We decided to explore some collaborations that involve not only scientists but artists to try to convey our messages about the importance and peril of the Tree of Life. We see that there is support for this approach in the recent American Academy report, Perceptions of Science in America. This report gives us some insight into how we might want to move forward. According to the report, although most Americans have a positive view of science and scientists, there are some lessons to be learned. For example, most Americans think that scientists should engage with policy-makers but they are also concerned about the use of science in public policy. The report found that confidence in scientific leaders varies based on demographics and on other factors. There are differences by race, gender, age, region of the country, educational background, and political party affiliation. And this means that as scientists, we need to find multiple ways to convey our message to a diverse public. Too often we engage with facts, and we fail to connect. More effective communication may be obtained through art. Science influences our minds, but art touches our hearts.
Our first foray into this art-science relationship was a collaboration with the University of Florida’s Harn Museum of Art. Selected works featuring biodiversity from the Harn’s collection were connected spatially and thematically by the Tree of Life throughout the museum. More recently, we developed the “One Tree, One Planet” project, which is still ongoing. “One Tree, One Planet” is an artistic project developed by Parisian artist, architect, and environmentalist Naziha Mestaoui. The work is a series of fifteen vignettes about biodiversity, each on a different species and set to music based on DNA sequences that are shared among all life but with a twist for particular species. The goal of “One Tree, One Planet” is to illustrate the connections among all species in the Tree of Life, including our connections with the tree, and to develop further connections among people around the globe.
As the show progresses, these fifteen vignettes come and go randomly and they are associated with a member of the audience, who serves as the guinea pig for and the representative of the human species. We have seen members of the audience taking photographs of the projections and engaging with each other. This led us to decide that we needed a smartphone app to increase and enhance the participatory elements. The launch of the app was on April 8, 2019. Then the show will travel to the Atlanta Botanical Garden. We hope to have showings in New York and St. Louis around Earth Day 2020.
Our second major project is a fifteen-minute animated movie about the Tree of Life, developed in collaboration with the Digital Worlds Institute at the University of Florida. This movie, called TreeTender, has been translated and is available with Korean, Chinese, Spanish, and Portuguese subtitles, as well as closed captioning in English. We held the premiere at the Florida Museum in 2017. The audience exceeded our expectations. Many hundreds of people crowded into the room to see the movie.
[Editor’s note: the audience that gathered at the Academy watched the movie trailer.]
We think that this combination of science and art may be the way forward, or one way forward at least, in helping us to convey some of the magnitude, beauty, and majesty of the Tree of Life with the public. Certainly, the information on the numbers of possible ways that we can put species together into a tree and various other sorts of big data questions are really exciting to scientists. But they may not be the most important and effective ways to communicate with the general public. And so we are delighted about the possibility of working with other scientists and artists to continue to develop these new methods for sharing our excitement about the Tree of Life and the role the tree has in helping us to understand both our place in the world and how best to continue to promote conservation efforts.
I would like to conclude with a quotation from that great biodiversity scientist, Albert Einstein: “Look deep into nature and then you will understand everything better.” The work that we have described this evening is the product of many collaborations. Our science work on the Tree of Life has been sponsored certainly by our university, the University of Florida, but also by the National Science Foundation. Our science and art work has been supported by the Florida Museum, the University of Florida, and the Digital Worlds Institute. We thank everyone who has contributed to this effort.
Questions & Answers
Question
I was wondering where viruses fit into the Tree of Life and what you view as their importance?
Douglas Soltis
Viruses are not incorporated into the Tree of Life that we shared with you this evening. As you may know, there is some controversy as to whether viruses are considered living. But nonetheless, there are many studies using tree-building methods to study viruses, and I mentioned one example, the SARS epidemic. Tree studies are critical to investigating those epidemics. How do they build the flu vaccine that you get yearly that predicts what will happen in that given year? They build a tree of relationships. They take viral strains that go back for over a hundred years and they build a tree of relationships. They then try to predict what the likely evolution of the virus will be into the next year.
Question
About twenty years ago, some scientists developed three criteria for assigning conservation priorities: representativeness, uniqueness, and irreplaceability. How would you apply those three criteria to phylogenetic diversity?
Pamela Soltis
Let me say that we certainly would not argue that phylogenetic diversity alone can be used as a criterion for selecting regions for conservation. Phylogenetic diversity measures are often highly correlated with species-richness measures and in many cases, those two approaches might give you very similar ideas. On the other hand, it is clear that in some cases phylogenetic diversity might actually capture a different element. It might serve to bring in that uniqueness component a little bit better than just a species number. In addition, as we see when we look at the Florida map, there are areas like the Everglades, which have a unique ecosystem, and we wouldn’t want not to conserve those areas because of low phylogenetic diversity. So that argues for not using phylogenetic diversity alone. In addition, there is a region in Florida that has very low phylogenetic diversity overall, but a very high number of locally endemic species, and that is in the Apalachicola River area. Basically, we think that phylogenetic diversity should be just one tool among many that are used. None of this should be done on the basis of a single criterion. The more tools we have at our disposal, the better we are able to make good decisions.
Question
There has been a considerable amount of evidence of horizontal gene transfer in early organisms and also in the retroviral fragments that make up a good portion of our genome. How does that affect the structure of the Tree of Life as you have depicted it?
Douglas Soltis
I think it is important to realize that the history of life is much more complicated than what is depicted on a tree with simple branching patterns, with descendant species that diverge from an ancestor. Branches of the tree can come back together through hybridization or horizontal gene transfer. The bacterial part of the Tree of Life is often represented as a net of life because there is so much gene exchange and horizontal gene transfer. To be honest, we have not done a very good job of using technology to depict these complex branching patterns. But I think that is the next step.
Pamela Soltis
Let me add that we also know that a lot of species do not originate through bifurcation. They originate through hybridization, coupled with genome duplication. And this is something that we see throughout the history of the plant branch of the Tree of Life. It also traces back in our own ancestry to rounds of whole-genome duplication in the common ancestry of all vertebrates. And so the Tree of Life is really one of branching, returning, branching, and so on. This is why it is so difficult to visualize. It is certainly one of those big challenges for the computer science people who might be interested in this sort of issue.
Question
I wonder whether there is an effort to have a record both of progressive development and of extinction in the Tree of Life?
Douglas Soltis
We like to talk in terms of a tree of life and death because we want to incorporate fossils into our giant trees. There are some versions of the tree that have fossils in them, dinosaurs placed in the tree, for example. There are attempts to do that on a broad scale. But as the trees get larger and larger, it becomes more and more difficult to do that on a comprehensive basis. That is why these projects are team science. You need computer scientists working with paleobiologists and others to put these kinds of comprehensive trees together.
Question
I am curious if you have done any audience analysis so far to see if the kinds of activities that you are giving people to improve their understanding of biodiversity concepts, such as trees, is working?
Pamela Soltis
We have a bit of data so far, which are still being analyzed, so I can’t give you a comprehensive answer yet. We distributed some audience questionnaires and conducted surveys and interviews following the premiere of the “One Tree, One Planet” and the TreeTender events. Some of the questioning had to do with whether the public understands what we mean by the Tree of Life. The questions were targeted at very general concepts. Once the data are fully analyzed, we hope to see how well the public understands the whole concept of things being connected. I think tree thinking is a fundamental part of biology that ought to be taught alongside the periodic table. That means, of course, that our teachers need to learn it in order to be able to teach it well.
Douglas Soltis
Let me add an interesting observation. We live in Florida and see people who do not necessarily believe in evolution. They watch TreeTender, which is basically teaching evolution without mentioning the word. And it seems they have no trouble with that. Why can we use the Tree of Life as a metaphor and it doesn’t bother them? They understand that we are all connected. Yet at the same time they have trouble with evolution.